#22: Spatial genetics for plant-based communities - and much more!
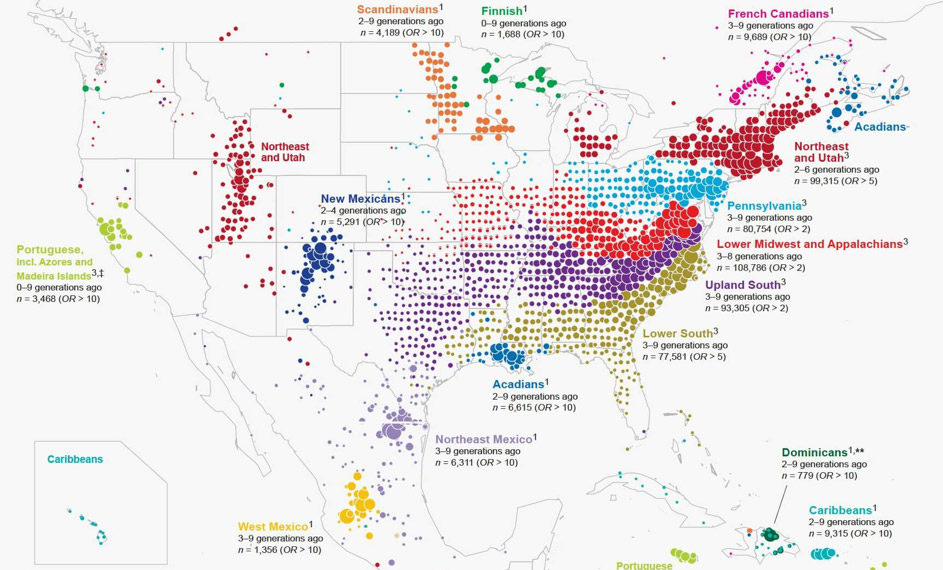
You might be familiar with an increasingly common application of spatial genetics research: using people's DNA to identify their ancestry, for example as done by 23andMe or ancestry.com. Variable pieces of DNA are used to identify geographically located clusters.
In our research, we take a more classical definition of genetics as heritable traits. Spatial genetics is thus the spatially resolved analysis of heritable traits, their frequency and distribution in communities, and how this affects adaptation and community interactions. Spatial genetics research includes approaches from ecology, remote sensing, chemistry, genetics, and bioinformatics.
Projects in spatial genetics use two basic approaches, analogous to forward and reverse genetics.
- Combine established remote sensing techniques with genetic and chemical analyses, and determine the correspondence between variation in remotely sensed data, and genetic variation (due to variation in specific aspects of biochemistry and structure).
- Determine traits of interest and design targeted remote sensing approaches.
Focussing on plant-based communities
Our research at GIUZ focuses on the spatial genetics of plant-based communities. This is because plants are the trophic basis of terrestrial ecosystems, and their diversity structures ecological communities. Specifically, plant genetic and species-level diversity have large effects. We need to understand the underlying mechanisms, and to employ them to achieve e.g. conservation or sustainability goals, and to improve our projections of future habitats under climate change. A spatially explicit understanding of genetics and chemical signaling in plant communities is needed. We furthermore require methods for large-scale and long-term monitoring. Remote sensing technologies can provide spatiotemporally resolved, large-scale, long-term data.
Laegern forest as a key research site
A key research site for us is the Laegern forest, which has been extensively mapped in terms of species and traits - and now, genotypes? Below is an image from work by Fabian Schneider, who mapped a part of the Laegern forest using a combination of laser scanning data for structure, and variation in imaging spectroscopy to identify differences among trees - but what causes those differences, and how much of it is genetic?
And by the way, spatially resolved genetic analyses also allow the visualization of the spread and evolution of the coronavirus HCoV-19.
Meredith C. Schuman